Post-process data in a trajectory
Stefan Widgren

Source: vignettes/post-process-data.Rmd
post-process-data.RmdAfter a model is created, a simulation is started with a call to the
run() function. The function returns a modified model
object containing a single stochastic solution trajectory. This
trajectory includes the state of each compartment recorded at every
time-point specified in tspan.
This vignette introduces the functionality in SimInf to
post-process and explore this trajectory data.
Let us first load the SimInf package.
Extract trajectory data with trajectory()
Most modeling studies require custom data analysis beyond simple
plotting. To support this, SimInf provides the
trajectory() method to extract the raw data as a
data.frame. This is useful if you need to:
- Perform custom statistical calculations (e.g., time to peak).
- Export data to CSV for use in other software.
- Combine results from multiple simulation runs.
Let’s simulate 10 days of data from an SIR model with 6 nodes. For reproducibility, we set the seed and specify the number of threads.
set.seed(123)
set_num_threads(1)
u0 <- data.frame(
S = c(100, 101, 102, 103, 104, 105),
I = c(1, 2, 3, 4, 5, 6),
R = c(0, 0, 0, 0, 0, 0)
)
model <- SIR(
u0 = u0,
tspan = 1:10,
beta = 0.16,
gamma = 0.077
)
result <- run(model)Extract the full trajectory data (all compartments, all nodes).
trajectory(result)## node time S I R
## 1 1 1 100 1 0
## 2 2 1 101 2 0
## 3 3 1 102 3 0
## 4 4 1 102 5 0
## 5 5 1 103 6 0
## 6 6 1 105 6 0
## 7 1 2 100 1 0
## 8 2 2 101 2 0
## 9 3 2 101 4 0
## 10 4 2 101 5 1
## 11 5 2 103 6 0
## 12 6 2 105 6 0
## 13 1 3 99 2 0
## 14 2 3 101 2 0
## 15 3 3 101 4 0
## 16 4 3 99 6 2
## 17 5 3 101 8 0
## 18 6 3 103 7 1
## 19 1 4 98 3 0
## 20 2 4 101 2 0
## 21 3 4 101 4 0
## 22 4 4 98 6 3
## 23 5 4 99 10 0
## 24 6 4 101 8 2
## 25 1 5 98 3 0
## 26 2 5 101 2 0
## 27 3 5 100 5 0
## 28 4 5 97 6 4
## 29 5 5 98 9 2
## 30 6 5 101 6 4
## 31 1 6 98 2 1
## 32 2 6 101 2 0
## 33 3 6 100 5 0
## 34 4 6 97 5 5
## 35 5 6 98 8 3
## 36 6 6 100 7 4
## 37 1 7 98 2 1
## 38 2 7 98 5 0
## 39 3 7 100 5 0
## 40 4 7 92 10 5
## 41 5 7 98 7 4
## 42 6 7 99 8 4
## 43 1 8 97 3 1
## 44 2 8 98 5 0
## 45 3 8 98 6 1
## 46 4 8 92 8 7
## 47 5 8 95 10 4
## 48 6 8 99 8 4
## 49 1 9 97 3 1
## 50 2 9 97 6 0
## 51 3 9 98 4 3
## 52 4 9 91 9 7
## 53 5 9 94 10 5
## 54 6 9 99 7 5
## 55 1 10 97 3 1
## 56 2 10 96 6 1
## 57 3 10 98 4 3
## 58 4 10 89 11 7
## 59 5 10 93 9 7
## 60 6 10 98 8 5Extract the number of recovered individuals (R) in the first node only.
trajectory(result, compartments = "R", index = 1)## node time R
## 1 1 1 0
## 2 1 2 0
## 3 1 3 0
## 4 1 4 0
## 5 1 5 0
## 6 1 6 1
## 7 1 7 1
## 8 1 8 1
## 9 1 9 1
## 10 1 10 1Extract the number of recovered individuals in the first and third nodes.
trajectory(result, compartments = "R", index = c(1, 3))## node time R
## 1 1 1 0
## 2 3 1 0
## 3 1 2 0
## 4 3 2 0
## 5 1 3 0
## 6 3 3 0
## 7 1 4 0
## 8 3 4 0
## 9 1 5 0
## 10 3 5 0
## 11 1 6 1
## 12 3 6 0
## 13 1 7 1
## 14 3 7 0
## 15 1 8 1
## 16 3 8 1
## 17 1 9 1
## 18 3 9 3
## 19 1 10 1
## 20 3 10 3Calculate prevalence from a trajectory using
prevalence()
The prevalence() function calculates the proportion of
individuals with the disease. It takes a model object and a formula:
- Left-hand side (LHS): Compartments representing “cases” (e.g., I).
- Right-hand side (RHS): Compartments representing the “at-risk” population (e.g., S + I + R).
The function also supports a level argument to change
the aggregation level:
-
level = 1(default): Prevalence aggregated over all nodes (global). -
level = 2: Proportion of nodes that have at least one case. -
level = 3: Prevalence calculated within each node (returns a matrix).
Let’s determine the proportion of infected individuals in the total population.
prevalence(result, I ~ S + I + R)## time prevalence
## 1 1 0.03616352
## 2 2 0.03773585
## 3 3 0.04559748
## 4 4 0.05188679
## 5 5 0.04874214
## 6 6 0.04559748
## 7 7 0.05817610
## 8 8 0.06289308
## 9 9 0.06132075
## 10 10 0.06446541Identical result is obtained with the shorthand I ~ .
(where . means “all compartments”).
prevalence(result, I ~ .)## time prevalence
## 1 1 0.03616352
## 2 2 0.03773585
## 3 3 0.04559748
## 4 4 0.05188679
## 5 5 0.04874214
## 6 6 0.04559748
## 7 7 0.05817610
## 8 8 0.06289308
## 9 9 0.06132075
## 10 10 0.06446541Calculate the proportion of nodes that are infected (at least one I individual).
prevalence(result, I ~ S + I + R, level = 2)## time prevalence
## 1 1 1
## 2 2 1
## 3 3 1
## 4 4 1
## 5 5 1
## 6 6 1
## 7 7 1
## 8 8 1
## 9 9 1
## 10 10 1Calculate the prevalence within each node individually.
prevalence(result, I ~ S + I + R, level = 3)## node time prevalence
## 1 1 1 0.00990099
## 2 2 1 0.01941748
## 3 3 1 0.02857143
## 4 4 1 0.04672897
## 5 5 1 0.05504587
## 6 6 1 0.05405405
## 7 1 2 0.00990099
## 8 2 2 0.01941748
## 9 3 2 0.03809524
## 10 4 2 0.04672897
## 11 5 2 0.05504587
## 12 6 2 0.05405405
## 13 1 3 0.01980198
## 14 2 3 0.01941748
## 15 3 3 0.03809524
## 16 4 3 0.05607477
## 17 5 3 0.07339450
## 18 6 3 0.06306306
## 19 1 4 0.02970297
## 20 2 4 0.01941748
## 21 3 4 0.03809524
## 22 4 4 0.05607477
## 23 5 4 0.09174312
## 24 6 4 0.07207207
## 25 1 5 0.02970297
## 26 2 5 0.01941748
## 27 3 5 0.04761905
## 28 4 5 0.05607477
## 29 5 5 0.08256881
## 30 6 5 0.05405405
## 31 1 6 0.01980198
## 32 2 6 0.01941748
## 33 3 6 0.04761905
## 34 4 6 0.04672897
## 35 5 6 0.07339450
## 36 6 6 0.06306306
## 37 1 7 0.01980198
## 38 2 7 0.04854369
## 39 3 7 0.04761905
## 40 4 7 0.09345794
## 41 5 7 0.06422018
## 42 6 7 0.07207207
## 43 1 8 0.02970297
## 44 2 8 0.04854369
## 45 3 8 0.05714286
## 46 4 8 0.07476636
## 47 5 8 0.09174312
## 48 6 8 0.07207207
## 49 1 9 0.02970297
## 50 2 9 0.05825243
## 51 3 9 0.03809524
## 52 4 9 0.08411215
## 53 5 9 0.09174312
## 54 6 9 0.06306306
## 55 1 10 0.02970297
## 56 2 10 0.05825243
## 57 3 10 0.03809524
## 58 4 10 0.10280374
## 59 5 10 0.08256881
## 60 6 10 0.07207207Visualize a trajectory with plot()
The plot() function provides a quick way to inspect the
outcome. It can display:
- The median and quantile range across all nodes.
- Individual trajectories for specific nodes.
- Prevalence curves.
Note: Since the simulation is stochastic, the exact lines shown below will vary unless set.seed() is used.
Aggregated View (Median and Range)
Plot the median and interquartile range (IQR) of all compartments.
plot(result)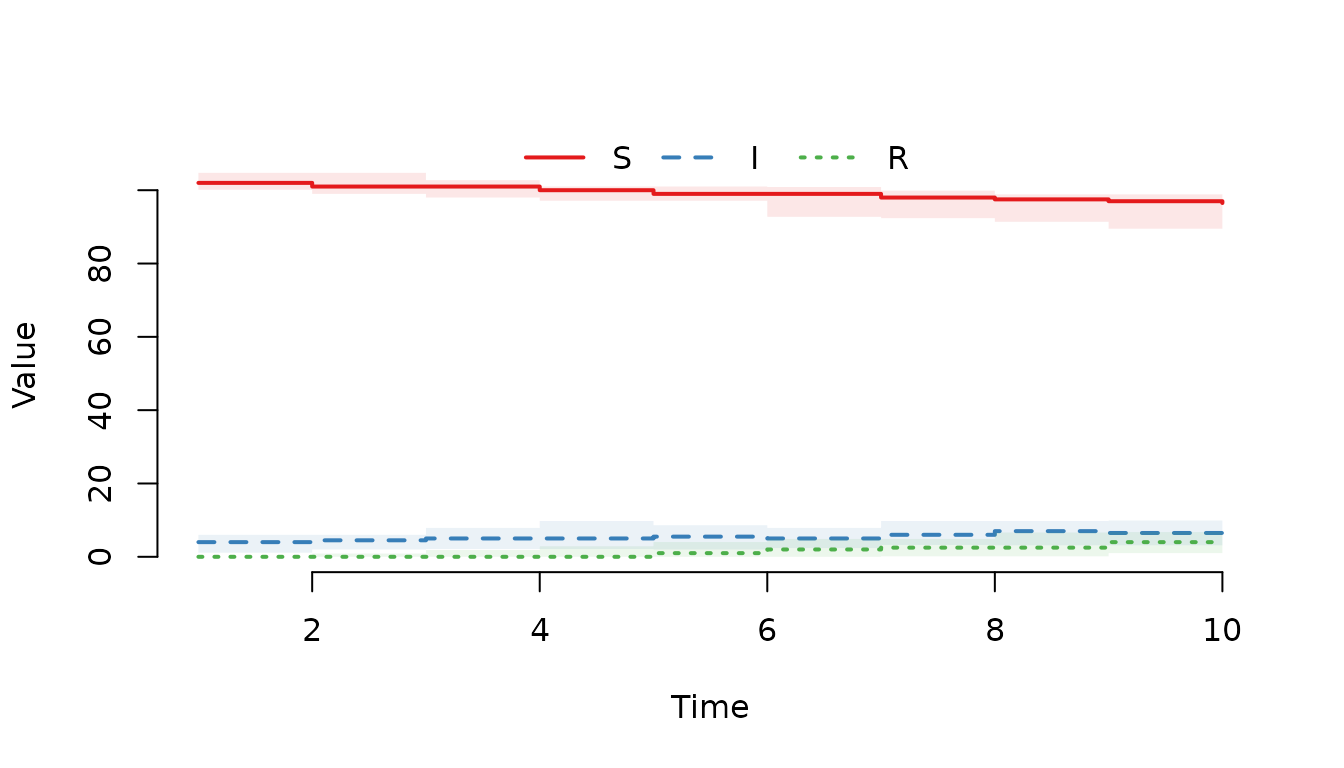
Plot the median and the middle 95% quantile range.
plot(result, range = 0.95)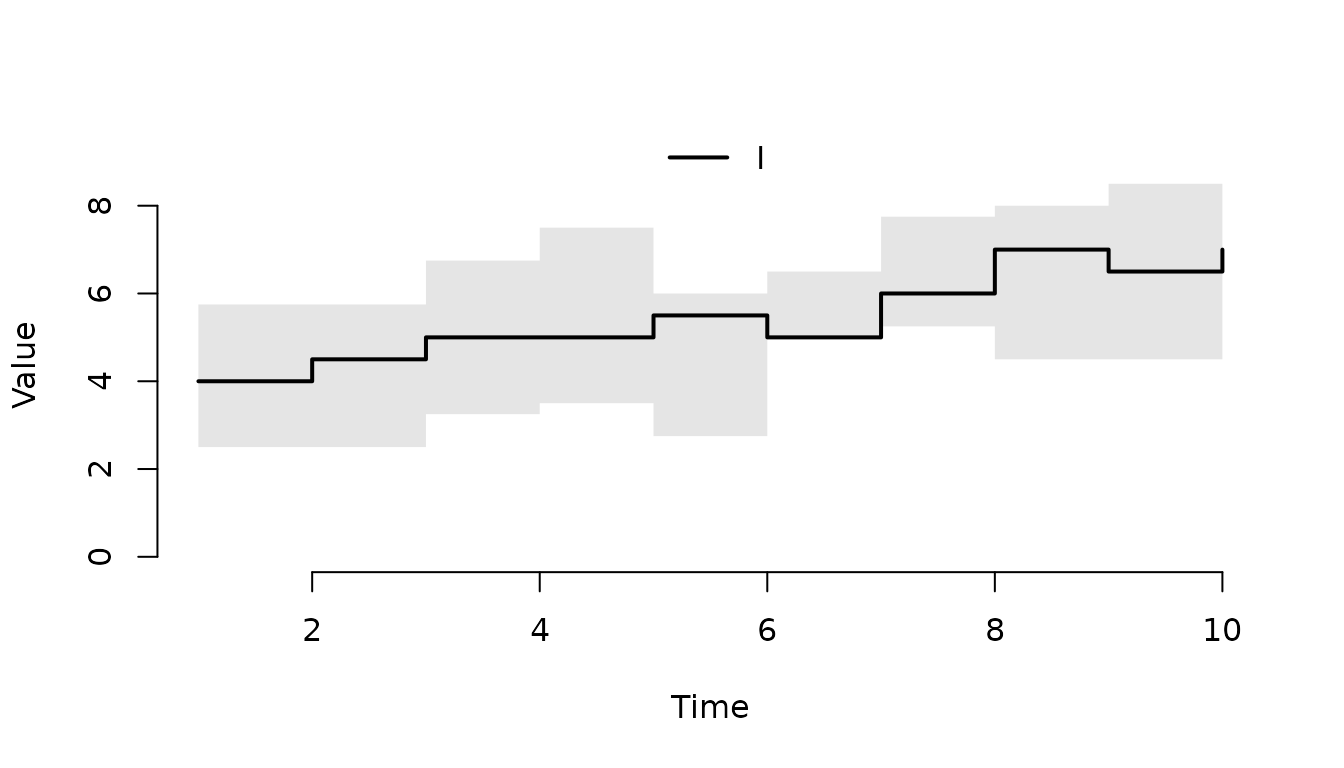
Plot only the infected individuals (I).
plot(result, "I")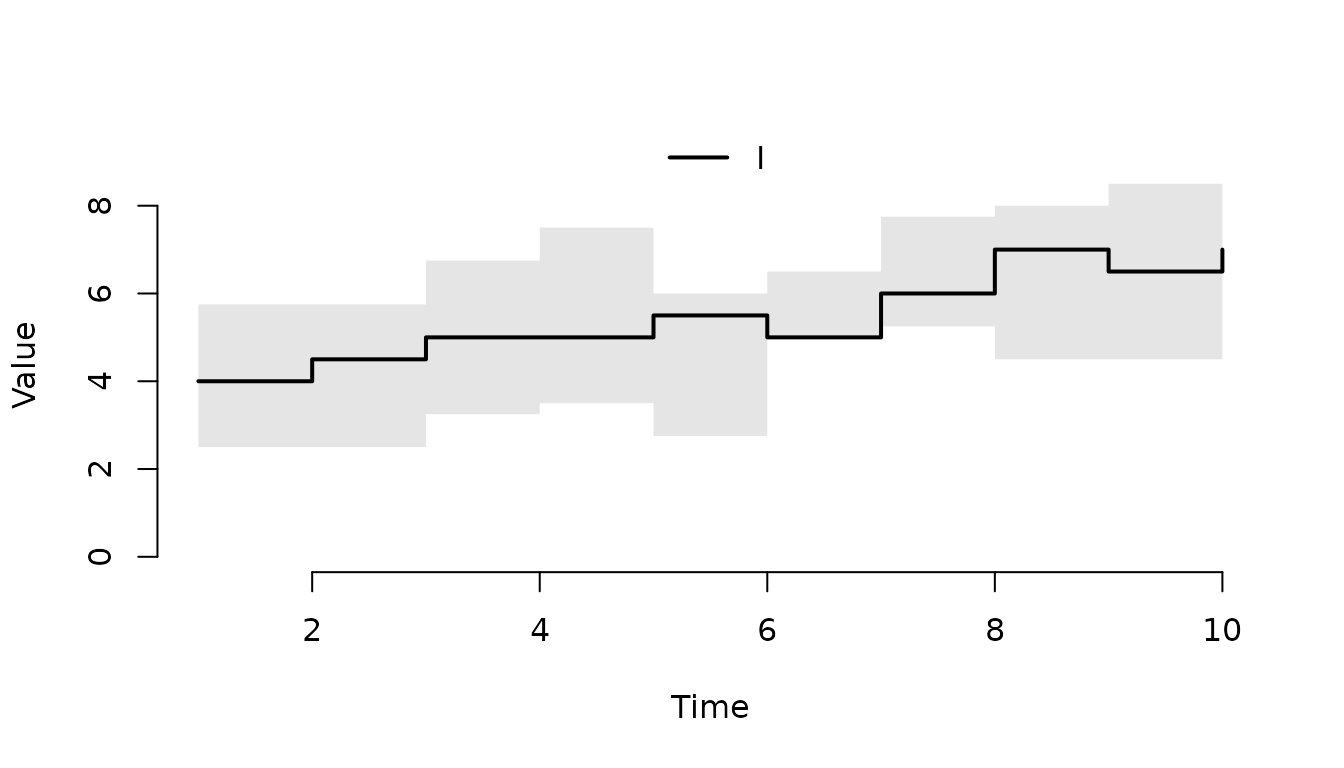
Use formula notation to plot the infected individuals.
plot(result, ~I)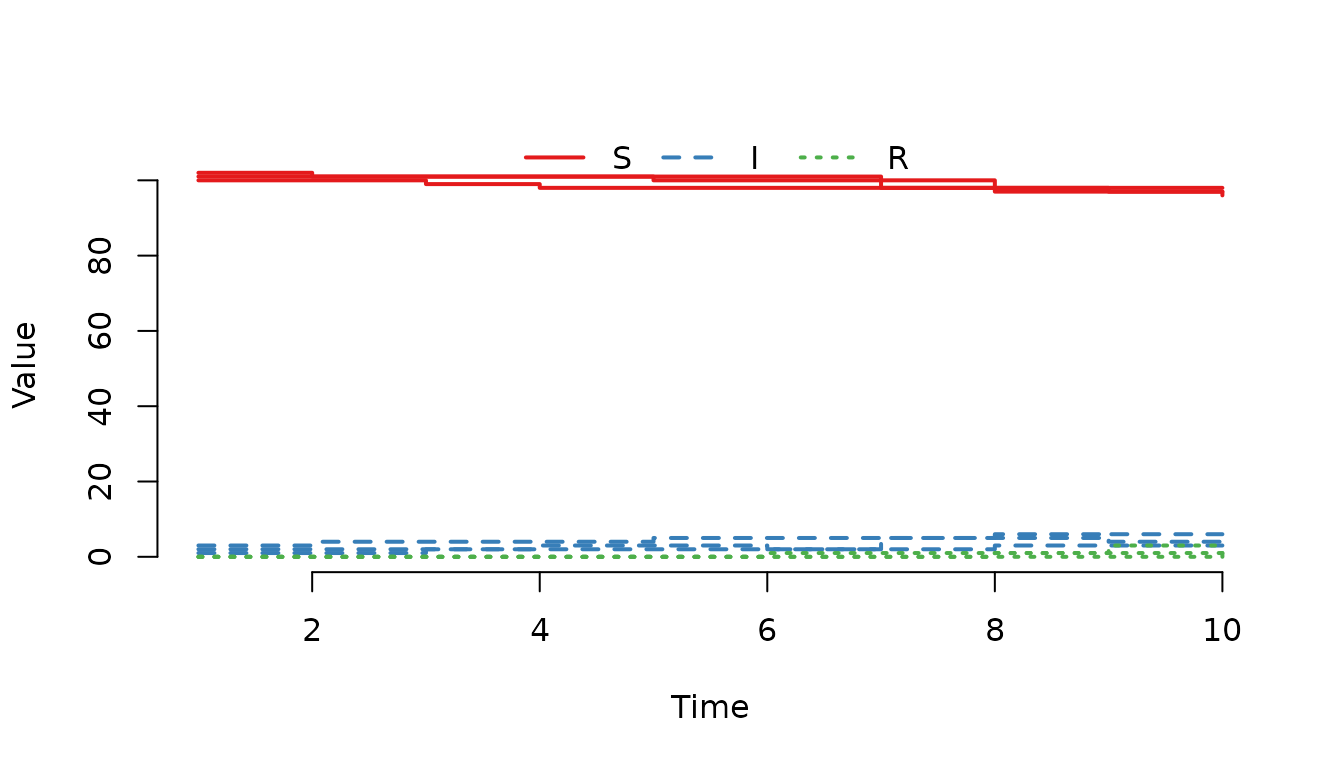
Individual Node View
Plot the trajectories for the first three nodes. We use
range = FALSE to suppress the shaded median/range bands and
show the individual lines.
plot(result, index = 1:3, range = FALSE)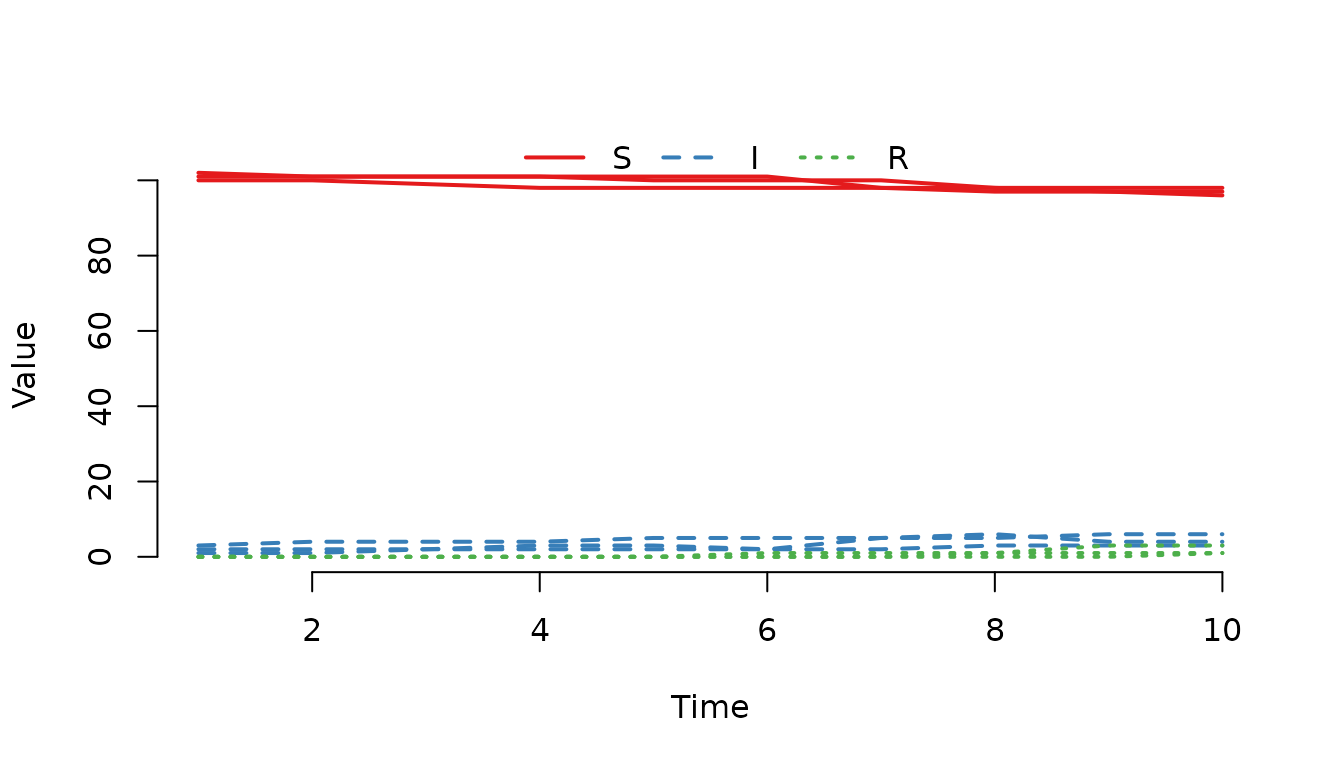
Use type = "l" to draw a line.
plot(result, index = 1:3, range = FALSE, type = "l")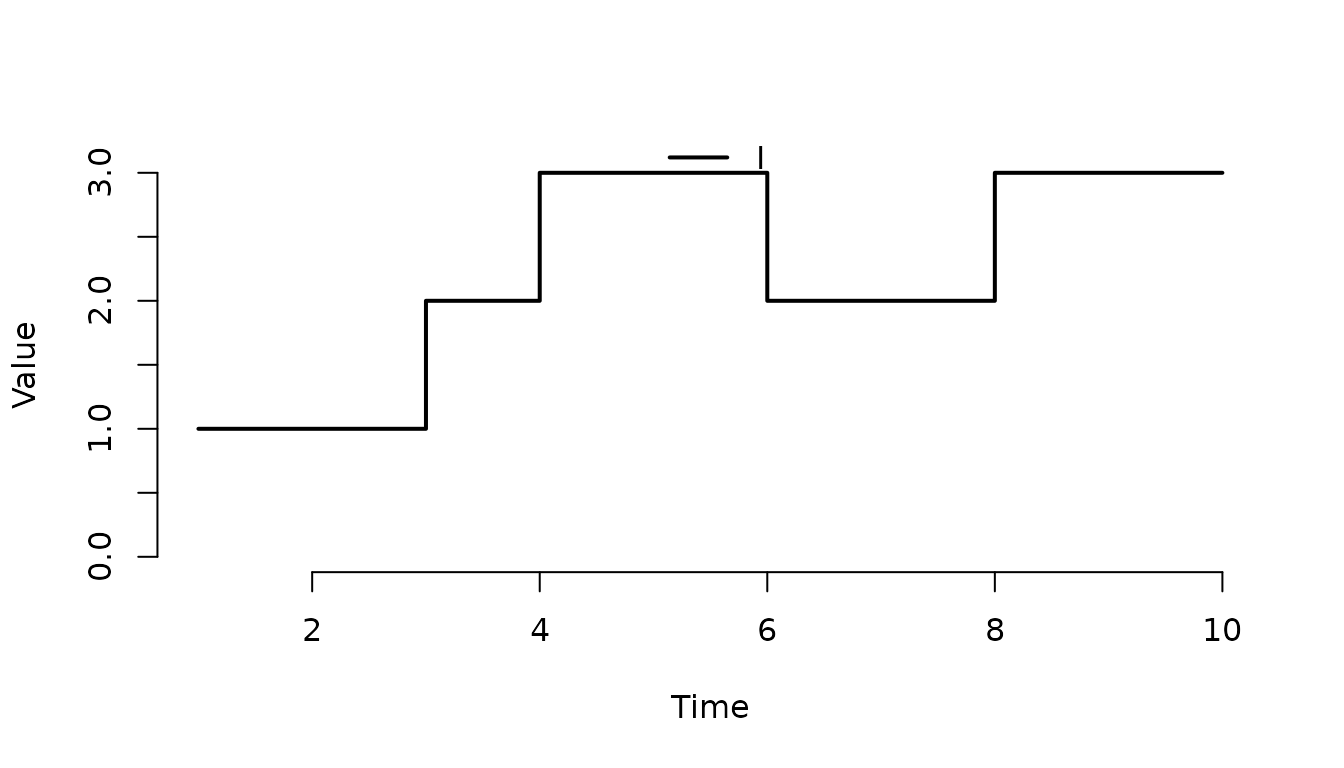
Plot the infected individuals in the first node only.
plot(result, "I", index = 1, range = FALSE)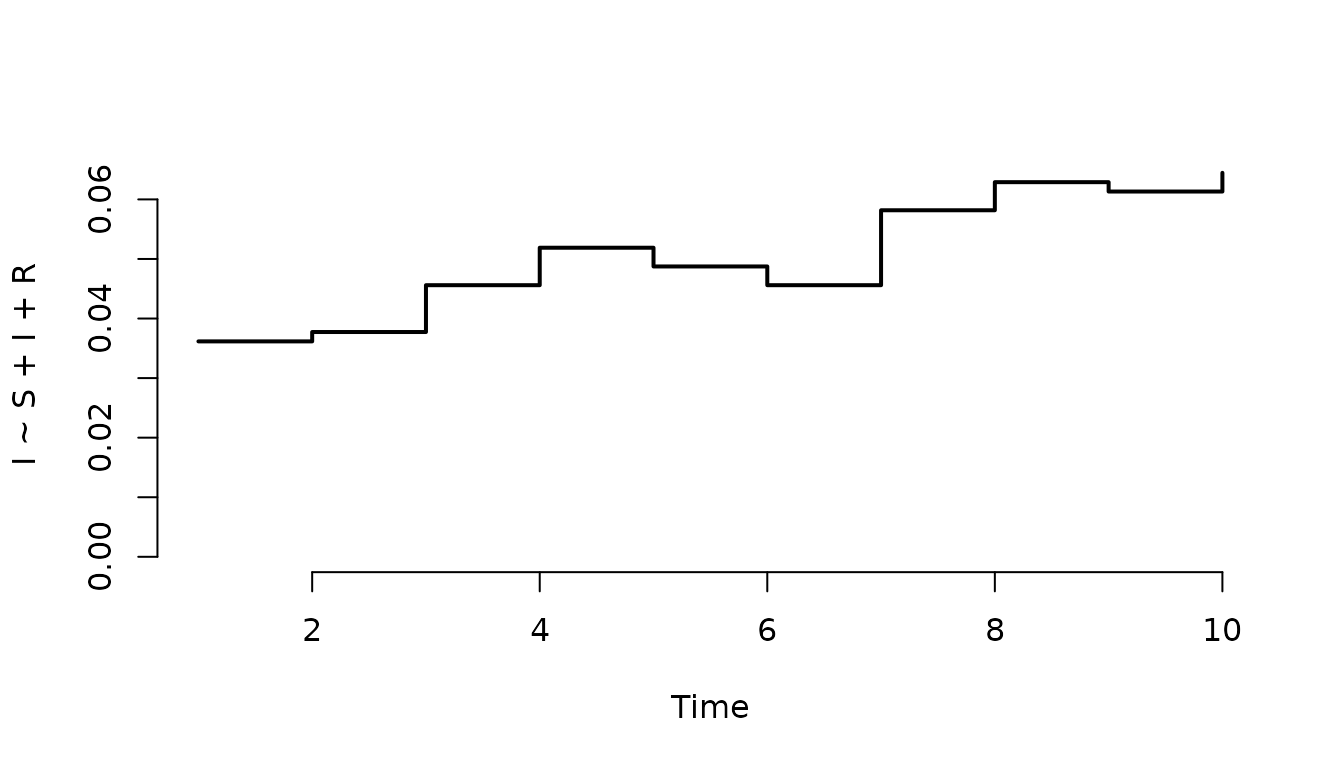
Prevalence Plots
Plot the proportion of infected individuals in the population.
plot(result, I ~ S + I + R)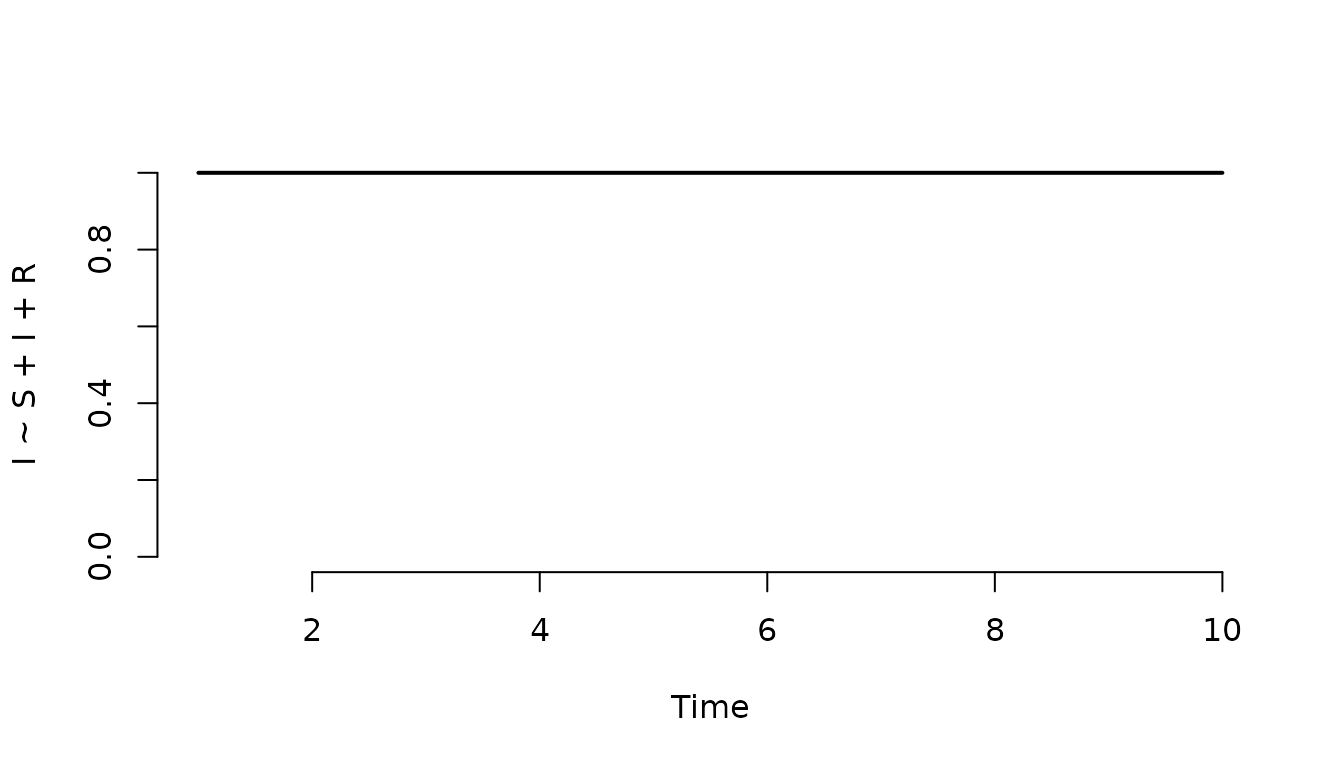
Plot the proportion of nodes with infected individuals
(level = 2).
plot(result, I ~ S + I + R, level = 2)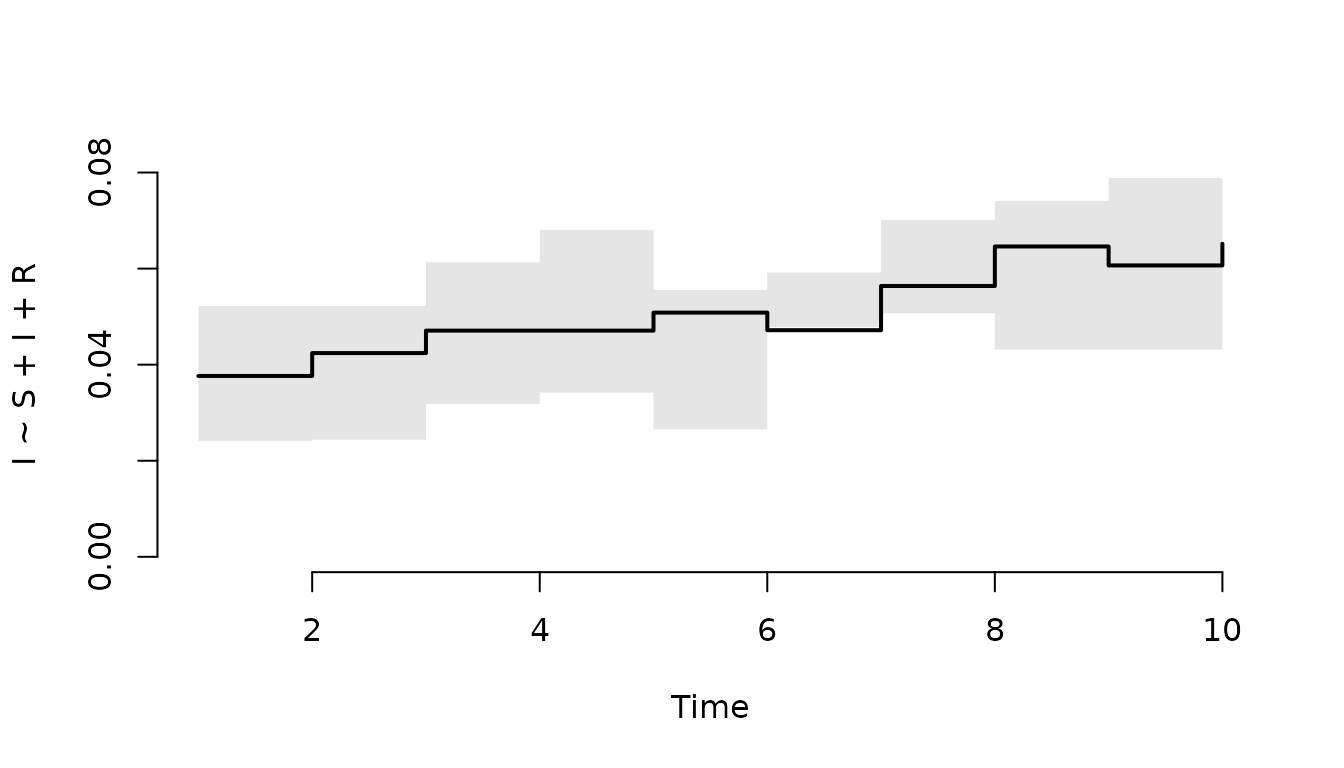
Plot the median and IQR of the prevalence within in each
node (level = 3).
plot(result, I ~ S + I + R, level = 3)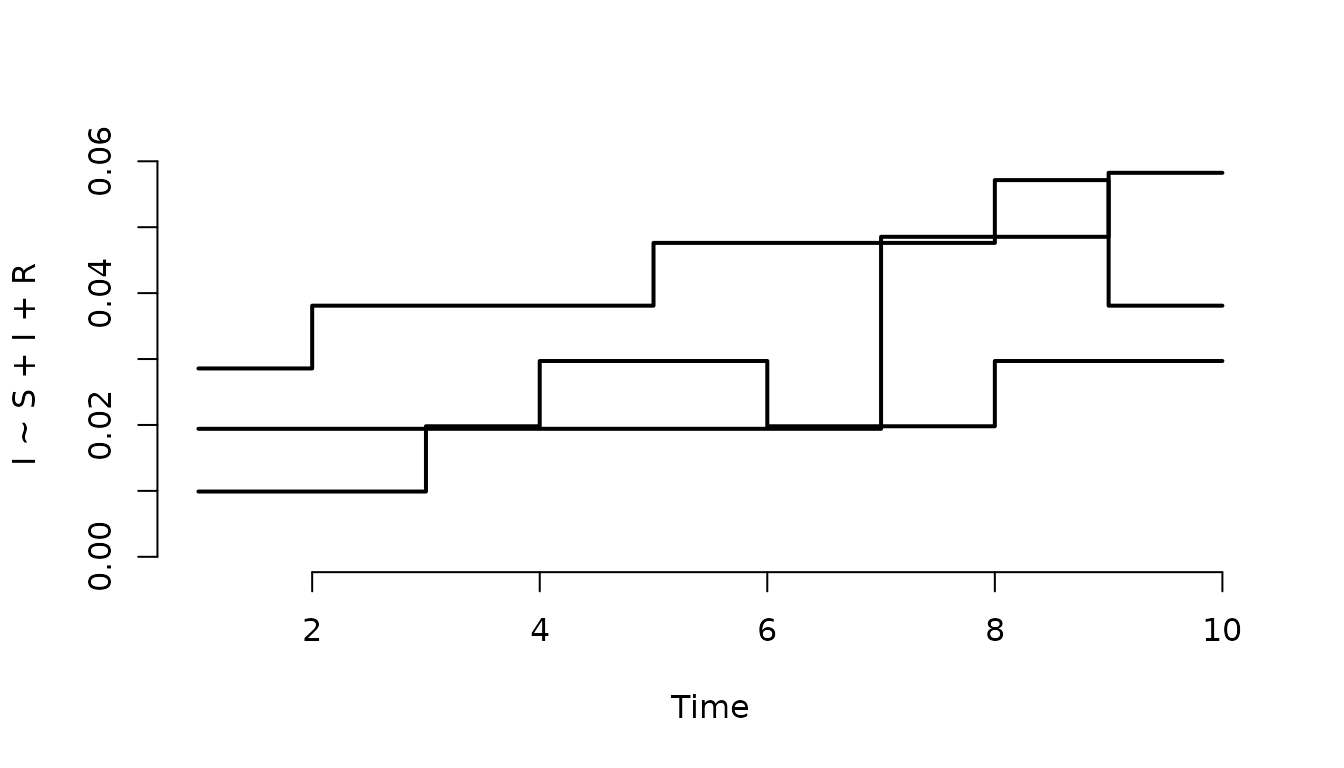
Plot the prevalence in the first three nodes.
plot(result, I ~ S + I + R, level = 3, index = 1:3, range = FALSE)
Summary
- Use
trajectory()to extract raw data for custom analysis. - Use
prevalence()to calculate disease proportions at different aggregation levels. - Use
plot()for quick visual inspection of medians, ranges, or individual trajectories.
To find more details on the plot method for SimInf_model
objects, run:
help("plot,SimInf_model-method", package = "SimInf")