Dataset containing 466,692 scheduled events for a population of 1,600 cattle herds over 1,460 days (4 years). Demonstrates how demographic and movement events affect SEIR dynamics in a cattle disease context.
Value
A data.frame with columns:
- event
Event type: "exit", "enter", or "extTrans".
- time
Day when event occurs (1-1460).
- node
Affected herd identifier (1-1600).
- dest
Destination herd for external transfer events.
- n
Number of cattle affected.
- proportion
0. Not used in this example.
- select
Model compartment to affect (see
SimInf_events).- shift
0. Not used in this example.
Details
The event data contains three types of scheduled events that affect cattle herds (nodes):
- Exit
Deaths or removal of cattle from a herd (n = 182,535). These events decrease the population and affect all disease compartments proportionally.
- Enter
Births or introduction of cattle to a herd (n = 182,685). These events add susceptible cattle to herds, increasing overall herd size.
- External transfer
Movement of cattle between herds (n = 101,472). These events transfer cattle from one herd to another, potentially facilitating between-herd disease transmission.
The select column in the returned data frame is mapped to
the columns of the internal select matrix:
select = 1corresponds to Enter events, targeting the Susceptible (S) compartment.select = 2corresponds to Exit and External Transfer events, targeting all compartments.
Events are distributed across all 1,600 herds over the 4-year period, reflecting realistic patterns of cattle demographic change and herd-to-herd movement. The timing and frequency of events can significantly influence disease dynamics simulated by the model.
See also
u0_SEIR for the corresponding initial cattle
population, SEIR for creating SEIR models with these
events, and SimInf_events for event structure
details
Examples
## For reproducibility, call the set.seed() function and specify the
## number of threads to use. To use all available threads, remove the
## set_num_threads() call.
set.seed(123)
set_num_threads(1)
## Create a 'SEIR' model with 1600 cattle herds (nodes) and initialize
## it to run over 4*365 days. Add ten exposed animals to the first
## herd. Define 'tspan' to record the state of the system at weekly
## time-points. Load scheduled events for the population of nodes with
## births, deaths and between-node movements of individuals.
u0 <- u0_SEIR()
u0$E[1] <- 10
model <- SEIR(u0 = u0,
tspan = seq(from = 1, to = 4*365, by = 7),
events = events_SEIR(),
beta = 0.16,
epsilon = 0.25,
gamma = 0.01)
## Display the number of cattle affected by each event type per day.
plot(events(model))
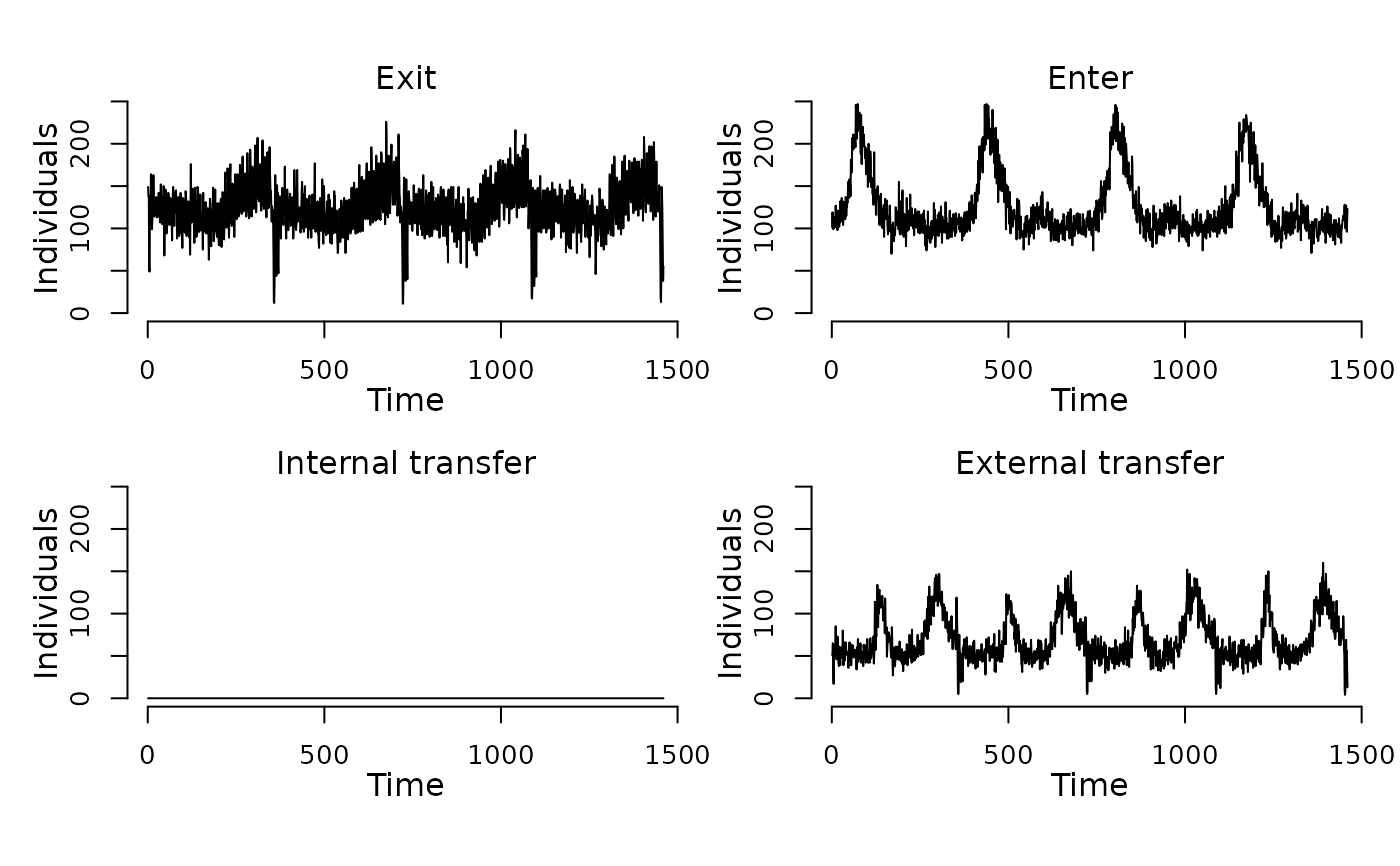 ## Run the model to generate a single stochastic trajectory.
result <- run(model)
## Plot the median and interquartile range of the number of
## susceptible, exposed, infected and recovered individuals.
plot(result)
## Run the model to generate a single stochastic trajectory.
result <- run(model)
## Plot the median and interquartile range of the number of
## susceptible, exposed, infected and recovered individuals.
plot(result)
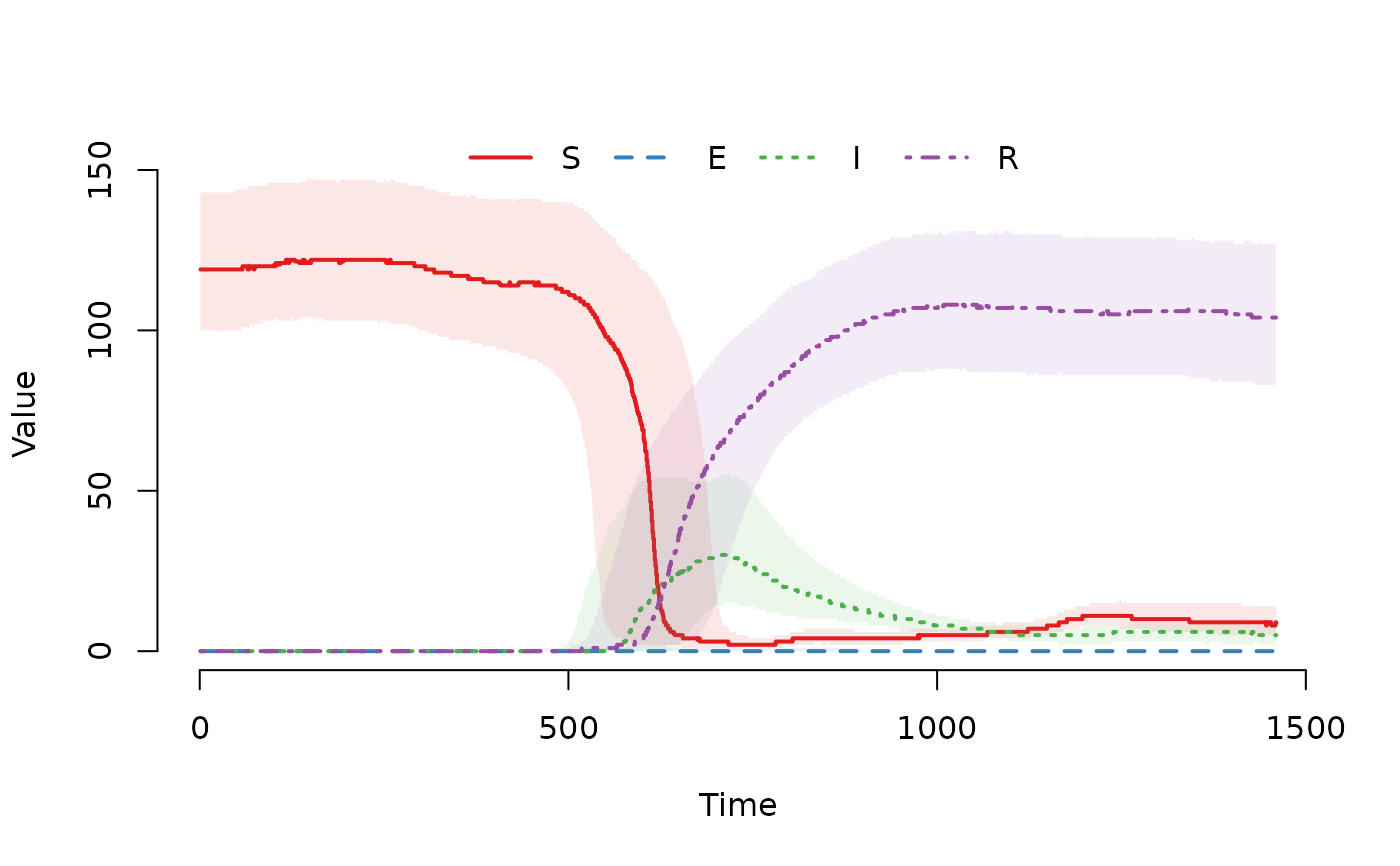 ## Plot the trajectory for the first herd.
plot(result, index = 1)
## Plot the trajectory for the first herd.
plot(result, index = 1)
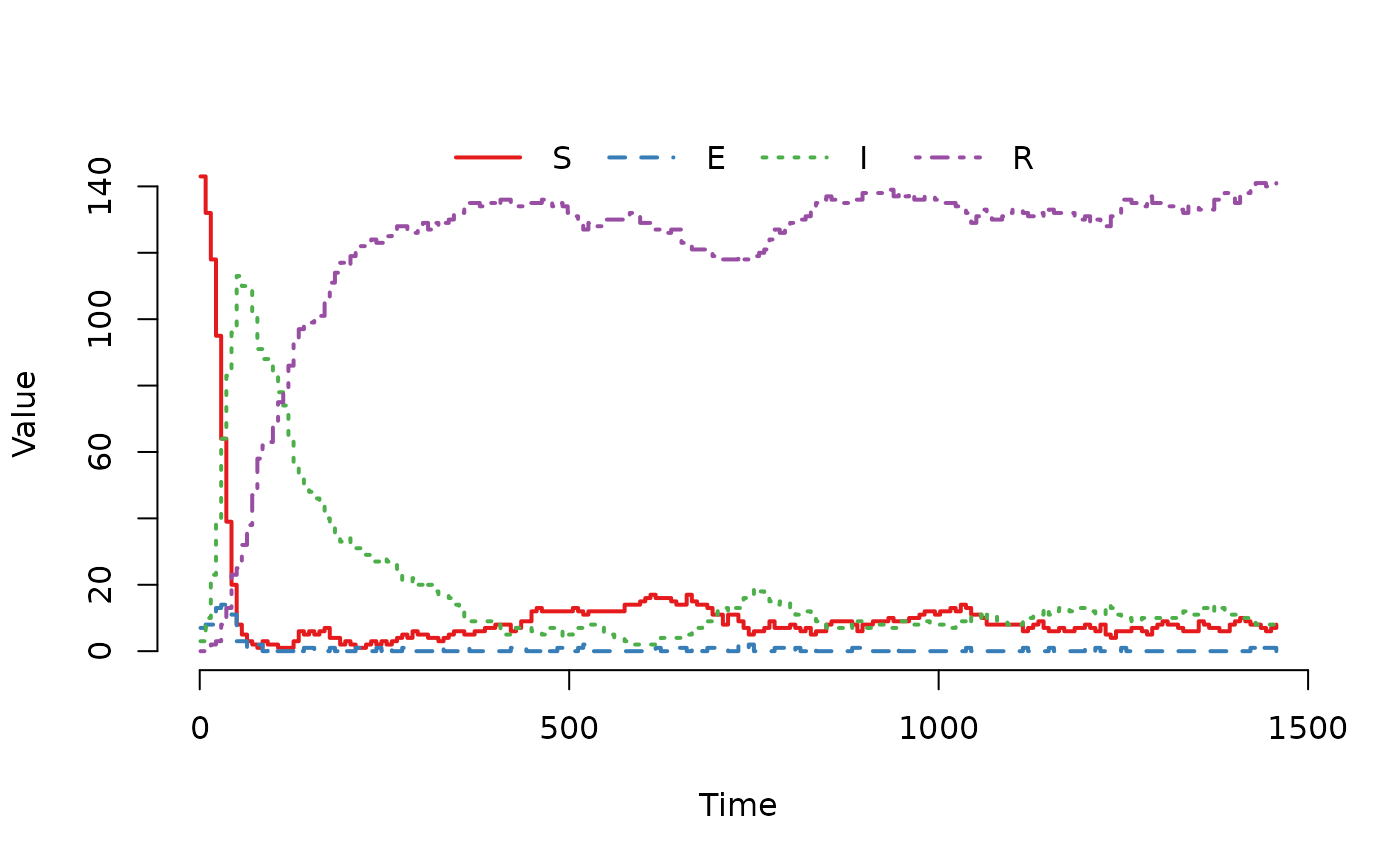 ## Summarize the trajectory. The summary includes the number of events
## by event type.
summary(result)
#> Model: SEIR
#> Number of nodes: 1600
#>
#> Transitions
#> -----------
#> S -> beta*S*I/(S+E+I+R) -> E
#> E -> epsilon*E -> I
#> I -> gamma*I -> R
#>
#> Global data
#> -----------
#> Number of parameters without a name: 0
#> - None
#>
#> Local data
#> ----------
#> Parameter Value
#> beta 0.16
#> epsilon 0.25
#> gamma 0.01
#>
#> Scheduled events
#> ----------------
#> Exit: 182535
#> Enter: 182685
#> Internal transfer: 0
#> External transfer: 101472
#>
#> Network summary
#> ---------------
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> Indegree: 40.0 57.0 62.0 62.1 68.0 90.0
#> Outdegree: 36.0 57.0 62.0 62.1 67.0 89.0
#>
#> Compartments
#> ------------
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> S 0.000 5.000 16.000 59.852 116.000 217.000
#> E 0.000 0.000 0.000 0.466 0.000 35.000
#> I 0.000 0.000 3.000 10.628 11.000 161.000
#> R 0.000 0.000 47.000 53.588 101.000 214.000
## Summarize the trajectory. The summary includes the number of events
## by event type.
summary(result)
#> Model: SEIR
#> Number of nodes: 1600
#>
#> Transitions
#> -----------
#> S -> beta*S*I/(S+E+I+R) -> E
#> E -> epsilon*E -> I
#> I -> gamma*I -> R
#>
#> Global data
#> -----------
#> Number of parameters without a name: 0
#> - None
#>
#> Local data
#> ----------
#> Parameter Value
#> beta 0.16
#> epsilon 0.25
#> gamma 0.01
#>
#> Scheduled events
#> ----------------
#> Exit: 182535
#> Enter: 182685
#> Internal transfer: 0
#> External transfer: 101472
#>
#> Network summary
#> ---------------
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> Indegree: 40.0 57.0 62.0 62.1 68.0 90.0
#> Outdegree: 36.0 57.0 62.0 62.1 67.0 89.0
#>
#> Compartments
#> ------------
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> S 0.000 5.000 16.000 59.852 116.000 217.000
#> E 0.000 0.000 0.000 0.466 0.000 35.000
#> I 0.000 0.000 3.000 10.628 11.000 161.000
#> R 0.000 0.000 47.000 53.588 101.000 214.000